TEEHR and CIROH Advance Cloud-Based Hydrologic Model Evaluation into a New Era
CIROH is advancing hydrologic model evaluation into a new era. Led by a core team of developers and scientists at RTI, with testing and contributions from others across the consortium, we've built TEEHR — a system purpose-built for evaluating models at scale. Combining novel approaches to data analytics with cutting-edge open data infrastructure, TEEHR enables a truly complete picture of model and forecast performance across datasets, sites, historical time periods, and forecast horizons.
Why do we need TEEHR?
TEEHR is built on a fundamental question: “Which hydrologic model is better?”. At its simplest, this can seem trivial; the model simulation is paired with observations, a few performance metrics are calculated, and you're on your way to a common performance analysis. Things can start to get more complicated if want to go larger, dig deeper or ask more nuanced questions:
- What if we want to compare many models against each other?
- What if we want to analyze thousands of locations with 40-years of hourly timestep data at the continental scale?
- What if we want to interrogate the data with questions like:
- “How does performance during high-flow events compare to low-flow events?”
- “How does model performance relate to physical basin attributes?”
- “What's the uncertainty associated with the resulting metrics?”
- What if we want to make the data easily accessible to the hydrologic community to support both historical and near-real time analyses?
These are the challenges TEEHR is designed to address.
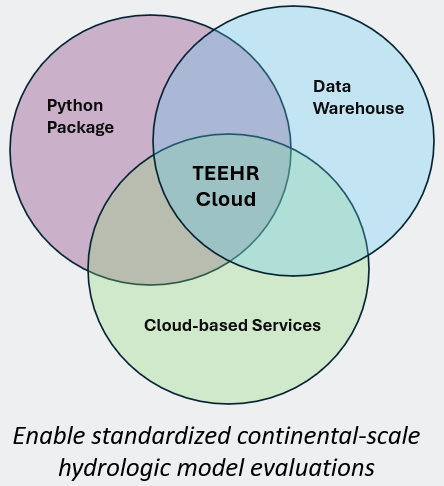
What is TEEHR?
The TEEHR Framework
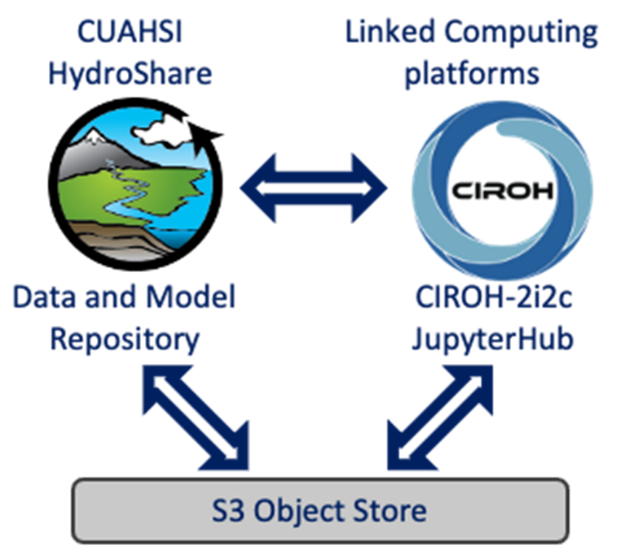
TEEHR began as a Python package enabling robust and iterative evaluation of hydrologic models. It has since expanded into a cloud-based platform (TEEHR-Cloud) consisting of a high-performance data warehouse optimized for huge analytic tables, a suite of cloud-based services supporting data exploration, automated processing and data ingests, and web-hosted interactive dashboards.
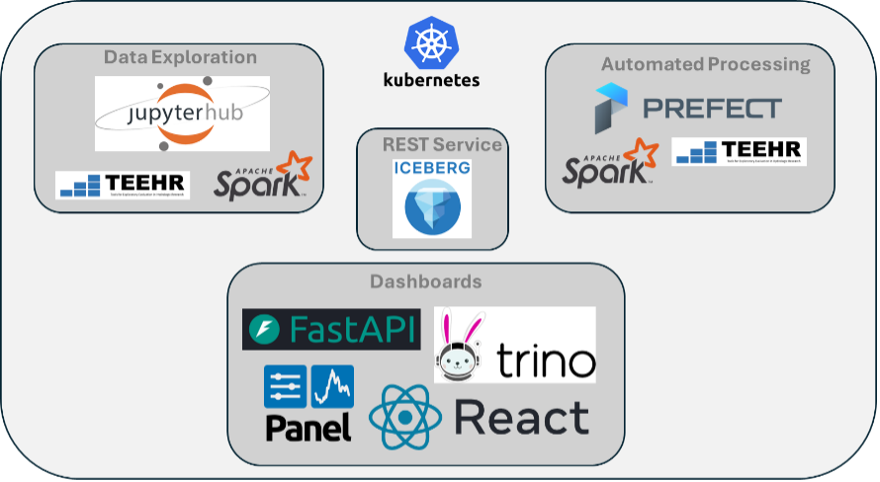
TEEHR-Cloud
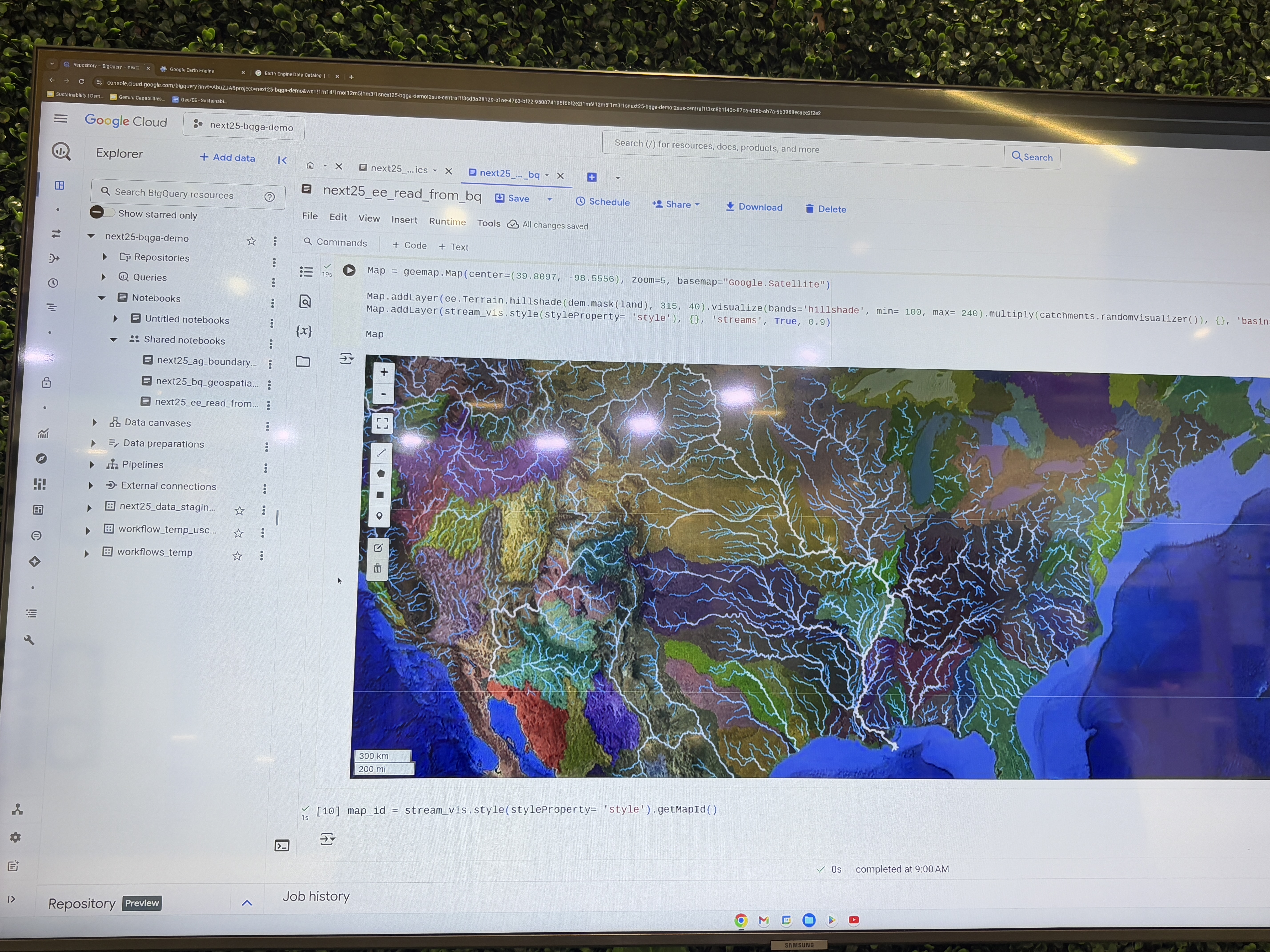
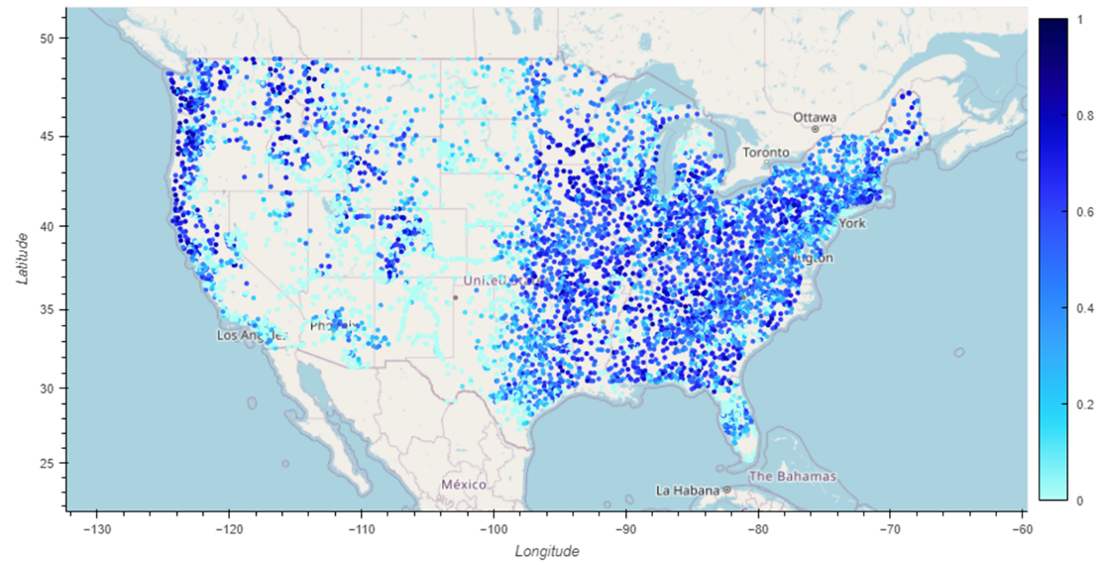
TEEHR-Cloud delivers evaluation-ready insights and datasets that are publicly available and continuously updated — including 40 years of hourly retrospective streamflow simulations across more than 8,000 gage sites, model forcings summarized to drainage basins (mean areal precipitation and temperature), and a real-time evaluation manager for live data ingestion and analysis of operational NWM, RFC forecasts and CIROH’s NextGen Research Data Stream (NRDS). It's all built on Apache Iceberg, an open table format designed for the demands of big data at scale and proven by well-known big data industries (e.g., Netflix).
TEEHR-Python
TEEHR-Python is the analytics engine that puts this data to work. Powered by PySpark for scalable, distributed compute and Apache Iceberg for robust data management — including time travel, upsert/append/delete operations, schema evolution, and ACID transactions — it gives analysts the full power of modern data engineering in a hydrologic context. An ever-growing suite of signature, deterministic, and probabilistic metrics, with sampling uncertainty estimation built-in, means users can interrogate model performance in ways that simply weren't easily accessible before. TEEHR-Python is still a Python package that users can install in their own environment via pip install teehr to use with their own data and to access datasets in the TEEHR-Cloud data warehouse.
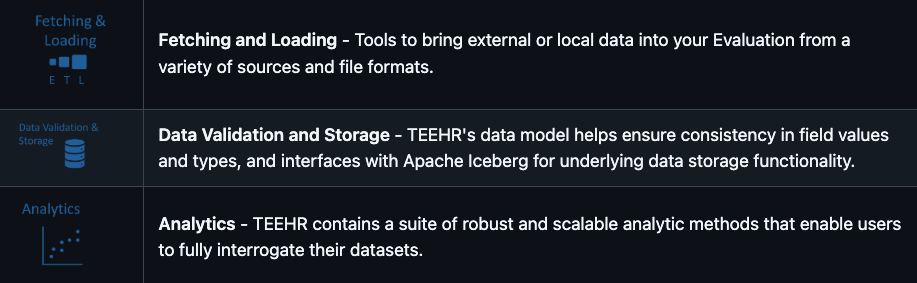

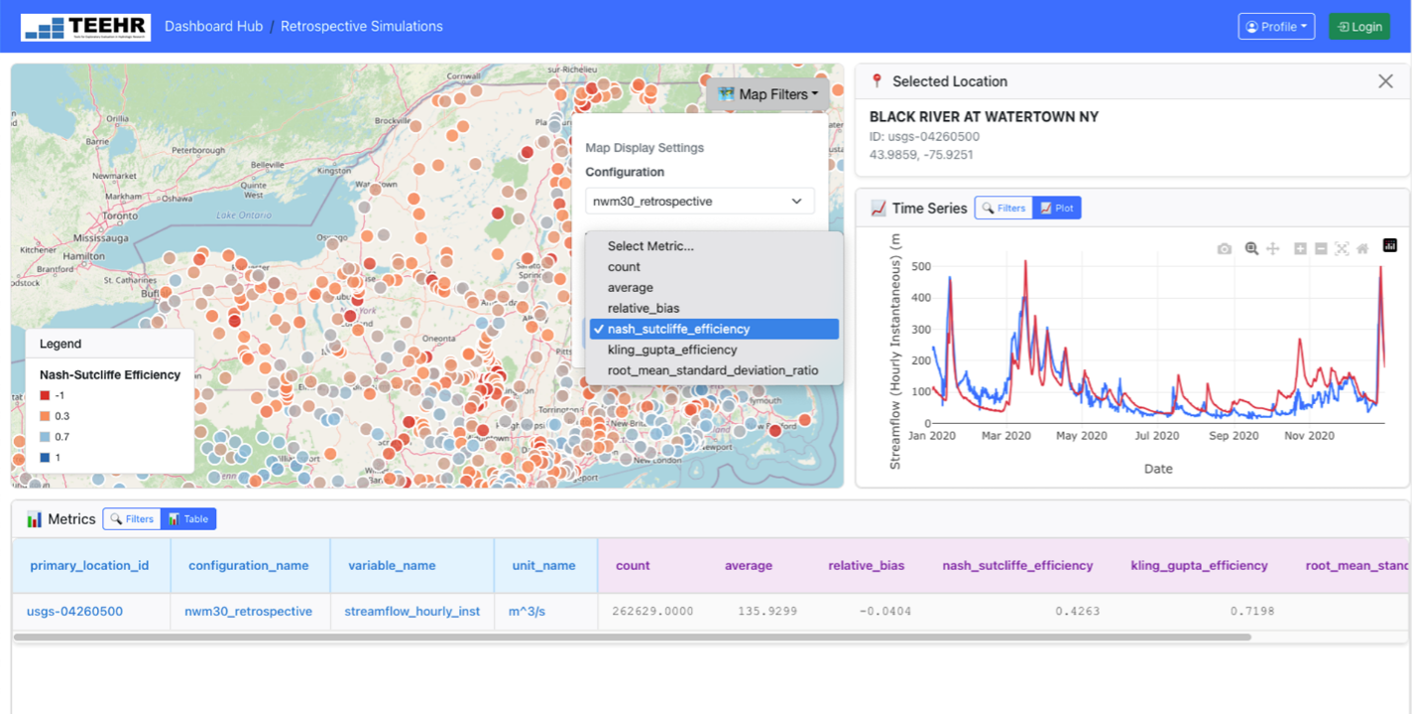
What are the TEEHR-Cloud Dashboards?
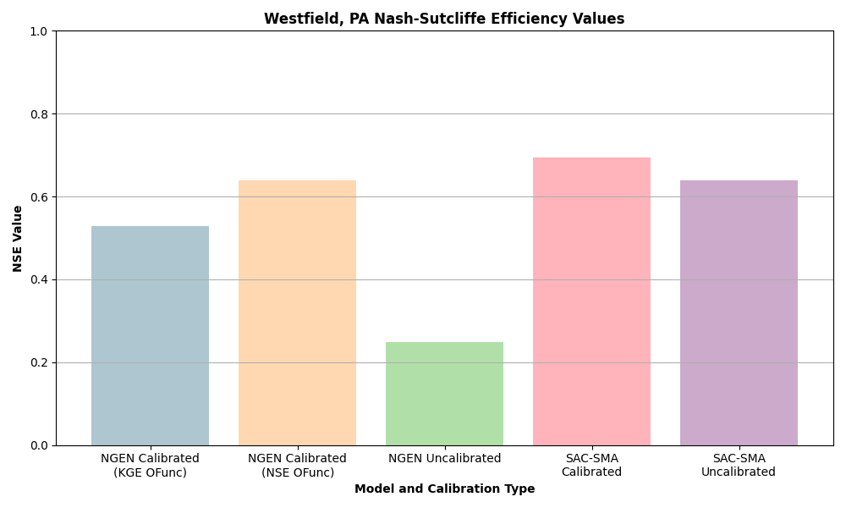
Several web-based dashboards, currently in development, allow users to view and interact with the data warehouse from application-specific perspectives. The two active dashboards include more general retrospective model comparisons and forecast analysis, while other, more application-specific, dashboards in development will focus on:
- Operational forecast (NWM, RFC) diagnostic and event-based evaluation
- Multi-model intercomparison to baselines with uncertainty
- Ensemble water supply forecasts
- Forecast Informed Reservoir Operations (FIRO)
- Flash flood forecasts
- Low flow forecasts
Summary
The TEEHR framework is under active development, as we continue to expand historical and real-time data ingests, application-specific dashboards, and core TEEHR-Python functionality. To learn more about TEEHR, the documentation is a good place to start (link below). If you want to ask a question, request a change or new feature, or report a bug, you can reach out through the CIROH #teehr slack channel, by creating an issue on the TEEHR GitHub repos, or by contacting us directly. Thanks for reading!
Additional Links:
- Documentation: https://hub.ciroh.org/docs/products/evaluation/rtiteehr/
- TEEHR documentation: https://rtiinternational.github.io/teehr/
- Dashboards (in-development): https://dashboards.teehr.rtiamanzi.org/
- TEEHR repo: https://github.com/RTIInternational/teehr
- TEEHR-Cloud repo: https://github.com/RTIInternational/teehr-hub
Our Team